Pointwise Transformations of Images
The function below allows us to perform a linear transformation on the image such that all pixels below a given threshold low_in are mapped to low_out and all pixels above high_in are mapped to high_out:
def image_contrast(pixel, low_in, high_in, low_out, high_out):
if pixel < low_in:
return low_out
elif pixel > high_in:
return high_out
else:
return low_out + (high_out - low_out)/(high_in - low_in)*(pixel - low_in)
f = np.vectorize(image_contrast)
For instance, the following set of values
low_in = 0.4, high_in = 0.6
low_out = 0.0, high_out = 1.0
corresponds to this transformation:
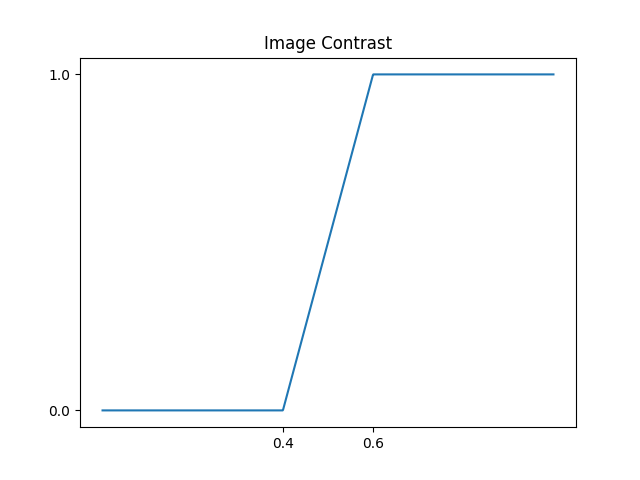
If we finally apply it to the camaraman image,
>>> camera = plt.imread('camera.png')
>>> camera_transform = f(camera, 0.4, 0.6, 0.0, 1.0)
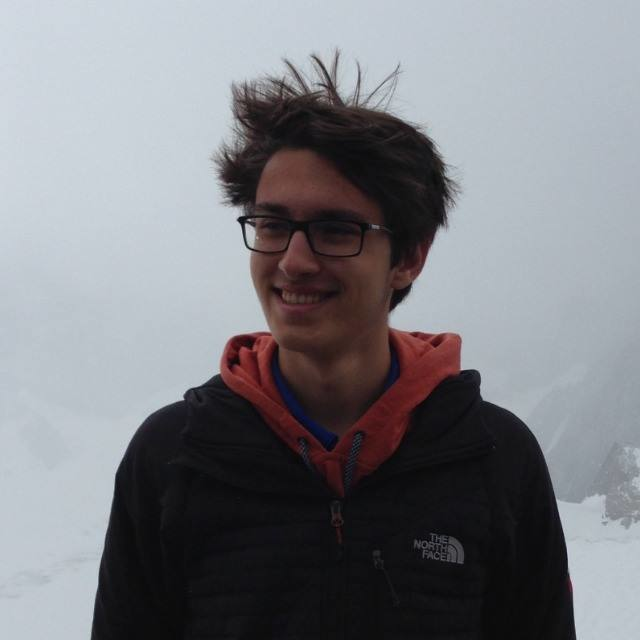
we observe the following effect:
``

Histograms and equalization
The following function allows us to visualize a list of images, together with its histogram and cumulative distribution. Moreover, it adds a cursor to each histogram that points out the value of each bin.
#!/usr/bin/env python3
import matplotlib.pyplot as plt
import numpy as np
from skimage import exposure
class SnaptoCursor(object):
def __init__(self, ax, x, y):
self.ax = ax
self.ly = ax.axvline(color='k', alpha=0.2) # the vert line
self.marker, = ax.plot([0],[0], marker="o", color="crimson", zorder=3)
self.x = x
self.y = y
self.txt = ax.text(0.7, 0.9, '')
def mouse_move(self, event):
if not event.inaxes:
return
x, y = event.xdata, event.ydata
indx = min(np.searchsorted(self.x, x), len(self.x) - 1)
x = self.x[indx]
y = self.y[indx]
self.ly.set_xdata(x)
self.marker.set_data([x],[y])
self.txt.set_text('x=%1.2f, y=%1.2f' % (x, y))
self.txt.set_position((x,y))
self.ax.figure.canvas.draw_idle()
def plot_histograms(images, nbins=256):
""" Plots each greyscale image in the list images,
together with its histogram and cumulative distribution
"""
n = len(images)
f, axes = plt.subplots(2, n)
axes = axes.reshape((2, n))
cursors_hist = [0 for i in range(n)]
# cursors_cdf = [0 for i in range(n)]
for i, image in enumerate(images):
ax_im, ax_hist = axes[:, i]
ax_cdf = ax_hist.twinx()
# Display image
ax_im.imshow(image, cmap='gray')
# Display histograms
hist, bins = exposure.histogram(image, nbins=256)
ax_hist.plot(bins, hist, color='black')
ax_hist.ticklabel_format(axis='y', style='scientific', scilimits=(0, 0))
ax_hist.set_xlabel('Pixel intensity')
if i == 0:
ax_hist.set_ylabel('Number of pixels')
# Display cumulative distribution
cdf, bins = exposure.cumulative_distribution(image, nbins=256)
ax_cdf.plot(bins, cdf, color='red')
ax_cdf.set_yticks([])
if i == len(images) - 1:
ax_cdf.set_ylabel('Fraction of total intensity')
ax_cdf.set_yticks(np.linspace(0, 1, 6))
# Display cursors
cursors_hist[i] = SnaptoCursor(ax_hist, bins, hist)
f.canvas.mpl_connect('motion_notify_event', cursors_hist[i].mouse_move)
# cursors_cdf[i] = SnaptoCursor(ax_cdf, bins, cdf)
# plt.connect('motion_notify_event', cursors_cdf[i].mouse_move)
plt.show()
return cursors_hist
In order to equalize a histogram we will use the equalize_hist() function from the skimage.exposure module:
>>> from skimage import exposure
>>> moon_equalized = exposure.equalize_hist(moon)
>>> moon_equalized
array([[0.74093608, 0.74093608, 0.94418847, ..., 0.0413489 , 0.04973926,
0.04973926],
[0.74093608, 0.74093608, 0.94418847, ..., 0.0413489 , 0.04973926,
0.04973926],
[0.74093608, 0.74093608, 0.94418847, ..., 0.0413489 , 0.04973926,
0.04973926],
...,
[0.23426186, 0.23426186, 0.44005065, ..., 0.787956 , 0.74093608,
0.74093608],
[0.5897308 , 0.5897308 , 0.52191412, ..., 0.83300147, 0.83300147,
0.83300147],
[0.5897308 , 0.5897308 , 0.52191412, ..., 0.83300147, 0.83300147,
0.83300147]])
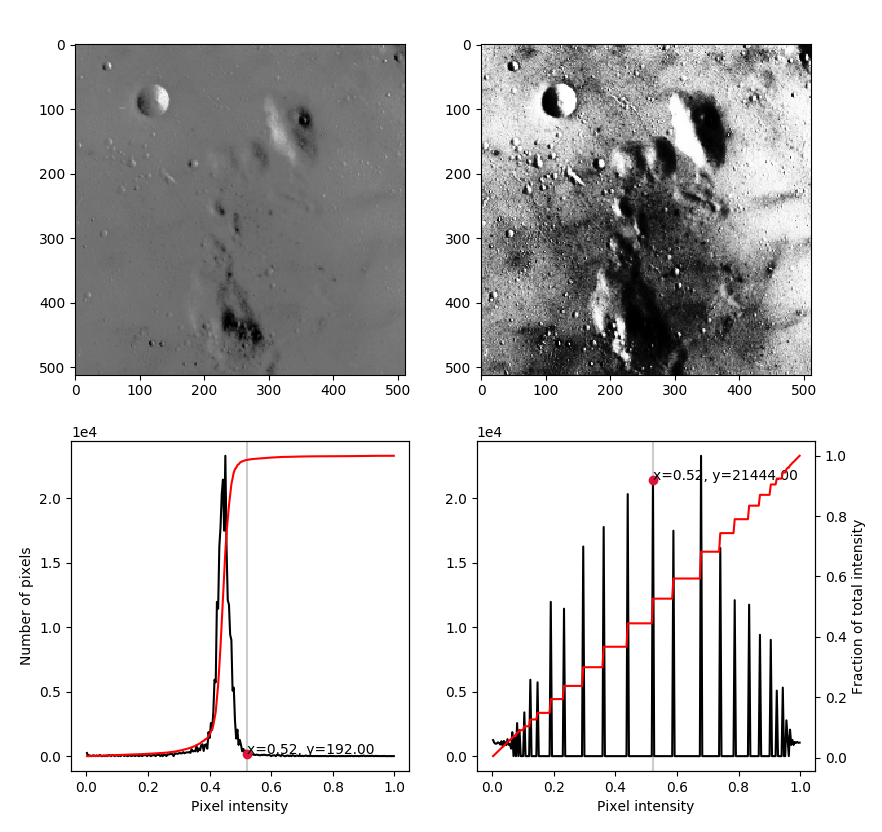
We can now use our previously defined function so see the effect of equalization on the image of the moon:
>>> images = [moon, moon_equalized]
>>> plot_histograms(images)
- Plotting with Spyder: In order to display figures in a separate window click Tools, Preferences, Ipython Console, Graphics and under Graphics Backend select automatic instead of inline.
- Plotting with Jupyter Notebooks: You need to execute the command
%matplotlib notebookfor interactive plots or%matplotlib inlinefor static images of your plot.
 Lliçons.jutge.org
Lliçons.jutge.org
Víctor Adell
Universitat Politècnica de Catalunya, 2023
Prohibit copiar. Tots els drets reservats.
No copy allowed. All rights reserved.